|
Quality science is supported by repetition. The consistency of an answer informs its reliability.
|
Approach to discovery-based research:
The Serbus group comprises research trainees ranging from high school interns to postdoctoral scholars. Lab members work in teams to tackle new research questions under the mentorship of senior lab members. Results that stand up to robust validation are regarded as new discoveries. The lab communicates findings to others in the form of oral presentations and research papers. The papers are submitted to academic journals, for assessment by colleagues who voluntarily serve as peer reviewers. Their insightful feedback contributes further to finalizing content of the manuscript. Publishing the work then shares the results with everyone. When possible, we aim to publish the work in an open-access format. This format has the advantage of making articles immediately accessible to the public for free. |
Current research questions:
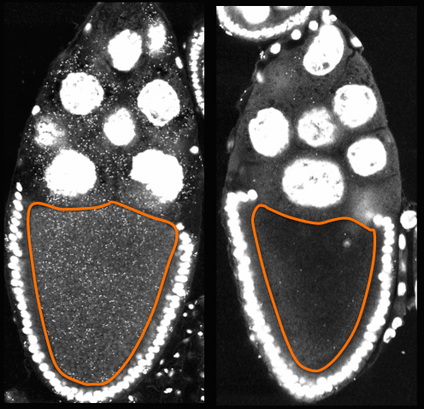
How is Wolbachia transmission achieved? Wolbachia spread maternally, via loading of the bacteria into developing eggs. Key requirements for transmission include 1) loading the egg with sufficient quantities of Wolbachia, and in some cases, 2) preferentially concentrating the Wolbachia at the site where future reproductive cells are specified. To understand how these outcomes are achieved, we apply genetic and chemical manipulations to fruit flies, then assess how their Wolbachia infections have responded. One of our primary approaches to analyze Wolbachia transmission is to fluorescently labeling Wolbachia in developing fly oocytes. The oocytes are then placed on glass slides and imaged by laser-scanning confocal microscopy. The images to the left show example data. Two fruit fly oocytes are shown, outlined in orange. These were dissected from Wolbachia-infected flies that ate regular food (left panel) and yeast-enriched food (right panel). You can see that the oocyte to the left is highly infected with Wolbachia (tiny dots), whereas the oocyte on the right carries very few Wolbachia. This comparison suggests that yeast-rich diets significantly affect Wolbachia colonization of oocytes. After many oocytes are imaged from multiple slide replicates, data from each condition are quantified and analyzed for statistical significance, which ultimately distinguish the findings of highest interest. |
Wolbachia colonization of fruit fly oocytes is nutrient-sensitive.
Slides imaged during winter break, 2015.
|
To date, we have found that intracellular transport, developmental patterning and nutritional signaling all contribute to maternal Wolbachia transmission. These cellular events have to date been considered mechanistically separate. With the support of the NSF, we are now investigating transmission as an integrated, functional process. Our work now includes mathematical modeling of oocyte colonization by Wolbachia, in collaboration with Distinguished Professor Michael Turelli at UC Davis.
|
We have also been developing complementary approaches to more broadly address the basis of Wolbachia density changes. One of these is quantitative real-time PCR. We are using this approach to analyze bodywide and tissue-specific Wolbachia densities in response to genetic, chemical and dietary treatments. |
|
What is basis for endosymbiont vitamin provisioning?
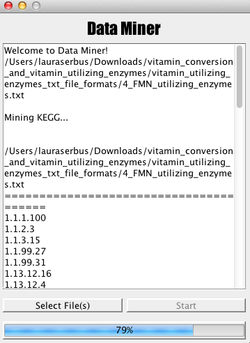
A new project in our lab has considered the nutritional basis for endosymbiosis in arthropods. Endosymbionts are widely described as provisioning vitamins to their hosts. To enable comparisons of vitamin production and vitamin use across endosymbiont taxa, we created a DataMiner tool that analyzes hundreds of vitamin-interacting enzymes. If this is also of interest to you, we welcome you to explore this resource. Create a .txt file with a list of enzymes (or use one of our comprehensive lists from HERE), listing one enzyme per line, described in terms of EC numbers such as “3.1.3.9”. Then upload the list into the Java version of DataMiner. It will cross-reference the enzyme list against 50 arthropod endosymbionts and 27 non-symbiont taxonomic relatives (described in terms of THESE abbreviations). When finished, DataMiner will deposit a tab-delimited set of results on your desktop. The Java-based DataMiner application (.jar) is HERE. The raw code files for DataMiner (.txt) are HERE and HERE. |
Preview of the DataMiner application, created by Brian Garcia Rodriguez.
|
If you wish to analyze a few (huge) lists of organisms that encode a given enzyme, to search for representation of specific organisms of interest, you are also welcome to try out the Microsoft Excel-based version of DataMiner (.xlsx) found HERE.